# A tibble: 94 × 3
Entity Year `Annual CO₂ emissions (per capita)`
<chr> <dbl> <dbl>
1 World 1750 0.0124
2 World 1760 0.0133
3 World 1770 0.0158
4 World 1780 0.0176
5 World 1790 0.0216
6 World 1800 0.0343
7 World 1810 0.0384
8 World 1820 0.0476
9 World 1830 0.0775
10 World 1840 0.0982
# ℹ 84 more rowsImport and Join Data
EE BIOL C177/C234
Today’s Menu 🎯
- Organize — RStudio Projects for tidy file structure
- Import —
read_csv()+janitor::clean_names() - Join — Combine data with
left_join() - Visualize — See the payoff of joining data!
Setup 📦
Packages for this chapter
File Organization
Use RStudio Projects!
- Keeps scripts, data, and output organized
- Sets the working directory automatically
- No more
setwd()nightmares! - File → New Project → New Directory
The here Package

Importing Data
Understanding File Paths 📍
When we render a .r file, the working directory is the folder containing that .r file.
my_project/
├── my_project.Rproj
├── data/
│ ├── per-capita-co-emissions.csv
│ └── temperature-anomaly.csv
└── slides/
└── 09_import_data.r <-- We are here!To reach data/, we must go up one level (../), then into data/:
"../data/per-capita-co-emissions.csv"
read_csv() — Read CSV Files
📥 Download per-capita-co-emissions.csv
Pro tip: Use the RStudio Import Dataset button to auto-generate code!
Clean Column Names
Messy column names → clean column names with one function:
# A tibble: 94 × 3
entity year annual_co2_emissions_per_capita
<chr> <dbl> <dbl>
1 World 1750 0.0124
2 World 1760 0.0133
3 World 1770 0.0158
4 World 1780 0.0176
5 World 1790 0.0216
6 World 1800 0.0343
7 World 1810 0.0384
8 World 1820 0.0476
9 World 1830 0.0775
10 World 1840 0.0982
# ℹ 84 more rowsAnnual CO2 Emissions Per Capita → annual_co2_emissions_per_capita ✨
Rename for Clarity
Chain it all together — clean, then rename:
data_co2 <- read_csv("../data/per-capita-co-emissions.csv") |>
janitor::clean_names() |>
rename(
co2 = annual_co2_emissions_per_capita
)
data_co2# A tibble: 94 × 3
entity year co2
<chr> <dbl> <dbl>
1 World 1750 0.0124
2 World 1760 0.0133
3 World 1770 0.0158
4 World 1780 0.0176
5 World 1790 0.0216
6 World 1800 0.0343
7 World 1810 0.0384
8 World 1820 0.0476
9 World 1830 0.0775
10 World 1840 0.0982
# ℹ 84 more rowsAlways Visualize After Import!
Sense-check: does the data look right?
data_co2 |>
ggplot(aes(x = year, y = co2)) +
geom_line() +
labs(
x = "Year",
y = "Annual CO2 emissions per capita"
) +
jtools::theme_nice()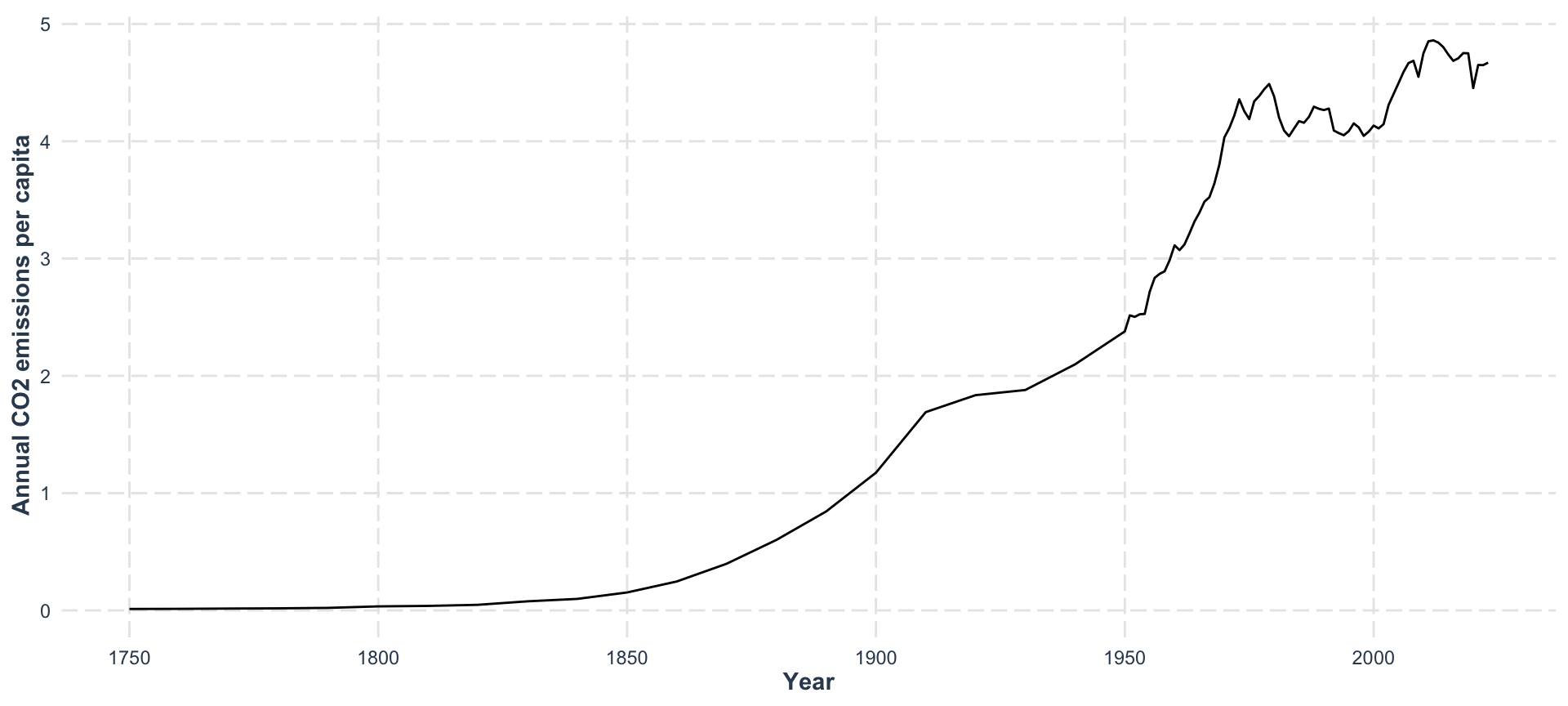
CO₂ is rising — but how does it relate to temperature? 🤔
Joining Data
Why Join?
Real data lives in multiple files
- CO₂ emissions data → one file
- Temperature data → another file
- Need to combine by a shared key (e.g.,
year)
Import the Second Dataset
📥 Download temperature-anomaly.csv
data_temp <- read_csv("../data/temperature-anomaly.csv") |>
janitor::clean_names() |>
select(year, global_average_temperature_anomaly_relative_to_1961_1990) |>
rename(temp_anomaly = global_average_temperature_anomaly_relative_to_1961_1990)
data_temp# A tibble: 175 × 2
year temp_anomaly
<dbl> <dbl>
1 1850 -0.418
2 1851 -0.233
3 1852 -0.229
4 1853 -0.270
5 1854 -0.292
6 1855 -0.297
7 1856 -0.320
8 1857 -0.467
9 1858 -0.389
10 1859 -0.281
# ℹ 165 more rowsNow we have two tibbles: data_co2 and data_temp
left_join() — The Workhorse
# A tibble: 94 × 4
entity year co2 temp_anomaly
<chr> <dbl> <dbl> <dbl>
1 World 1750 0.0124 NA
2 World 1760 0.0133 NA
3 World 1770 0.0158 NA
4 World 1780 0.0176 NA
5 World 1790 0.0216 NA
6 World 1800 0.0343 NA
7 World 1810 0.0384 NA
8 World 1820 0.0476 NA
9 World 1830 0.0775 NA
10 World 1840 0.0982 NA
# ℹ 84 more rowsKeeps all rows from the left, matches from the right
Join Types
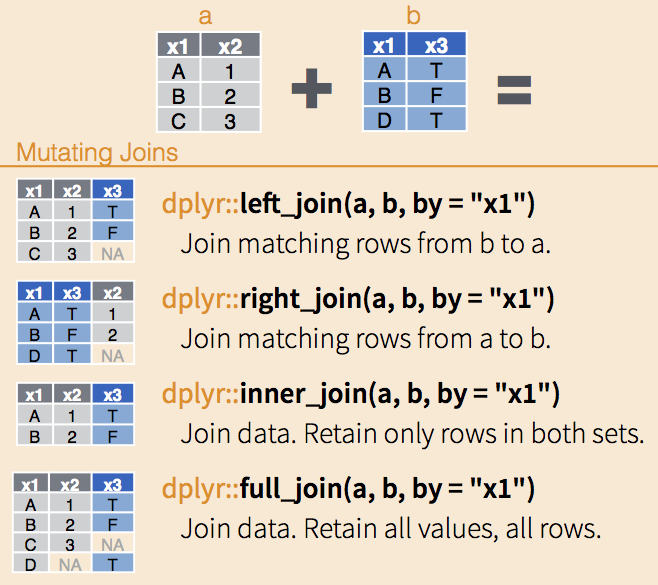
Join Types — Quick Reference
| Join | Keeps |
|---|---|
left_join() |
All rows from left |
right_join() |
All rows from right |
inner_join() |
Only matching rows |
full_join() |
All rows from both |
Best practice: Stick to left_join() for consistency
⚠️ Join Order Matters!
Joins are asymmetric — flipping changes the result!
The left side is your primary dataset — all its rows are always kept!
The Payoff: CO₂ vs Temperature 🌡️
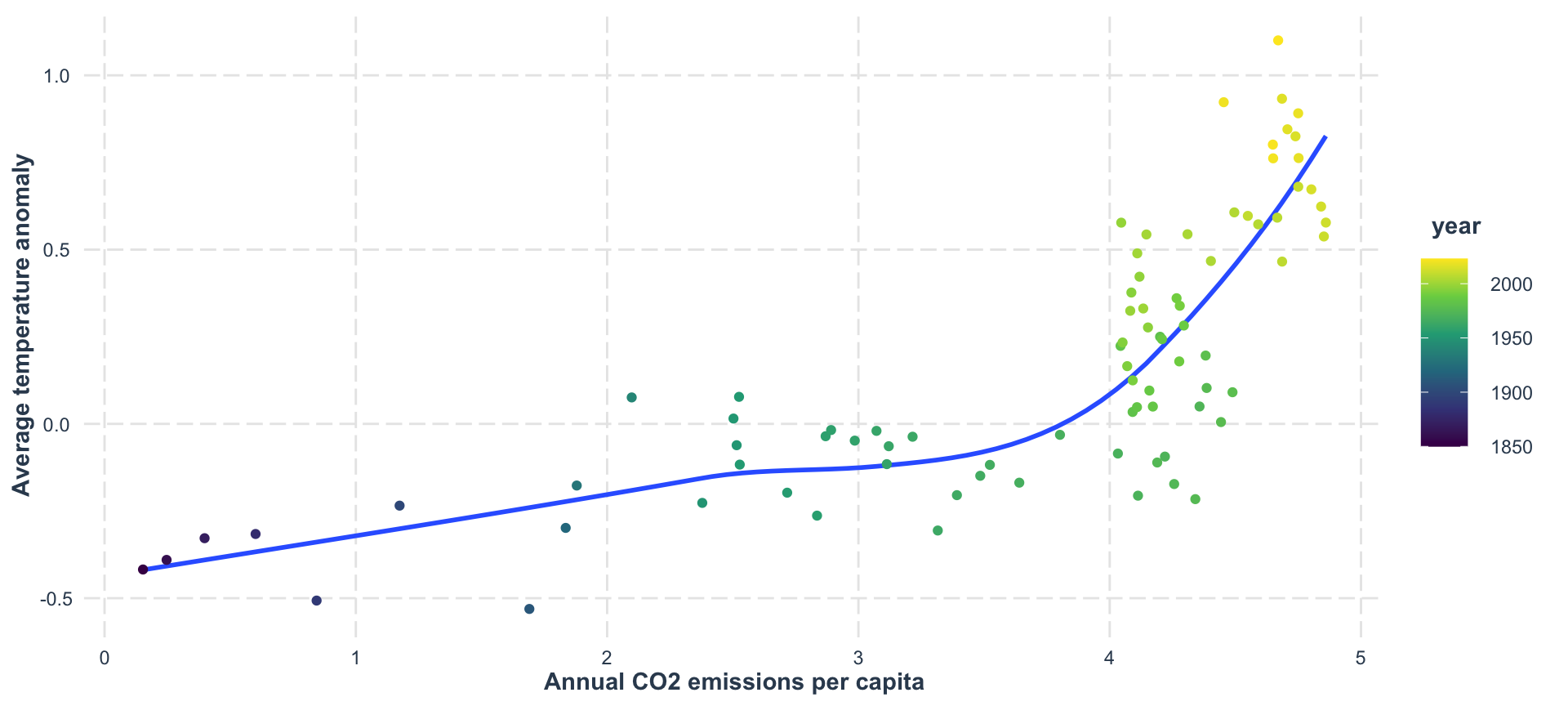
This is why we learn to join data — to answer real questions!
Summary
- Use RStudio Projects for organization
read_csv()+janitor::clean_names()for clean imports- Always visualize your data after importing!
left_join()combines data by shared keys- Join order matters — the left side is primary
- The real power: joining lets you answer new questions
Import and Join Data