# A tibble: 3 × 3
region `2007` `2009`
<chr> <dbl> <dbl>
1 Pacific 1039 2587
2 Southwest 51 176
3 Rocky Mountains and Plains 200 338Five Verbs for Tidying Data
EE BIOL C177/C234
Packages for this chapter
If you haven’t installed the following packages, run this code:
Today’s Menu 🎯
pivot_longer()— wide → longpivot_wider()— long → wideseparate()— split columnsdrop_na()/replace_na()— handle missing values- Tidy a real dataset — the Datasaurus Dozen 🦕
Tidy Data Reminder
Tidy datasets are all alike, but every messy dataset is messy in its own way. — Hadley Wickham
- Every row = one observation
- Every column = one variable
- Every cell = one value

Why Tidy Data?
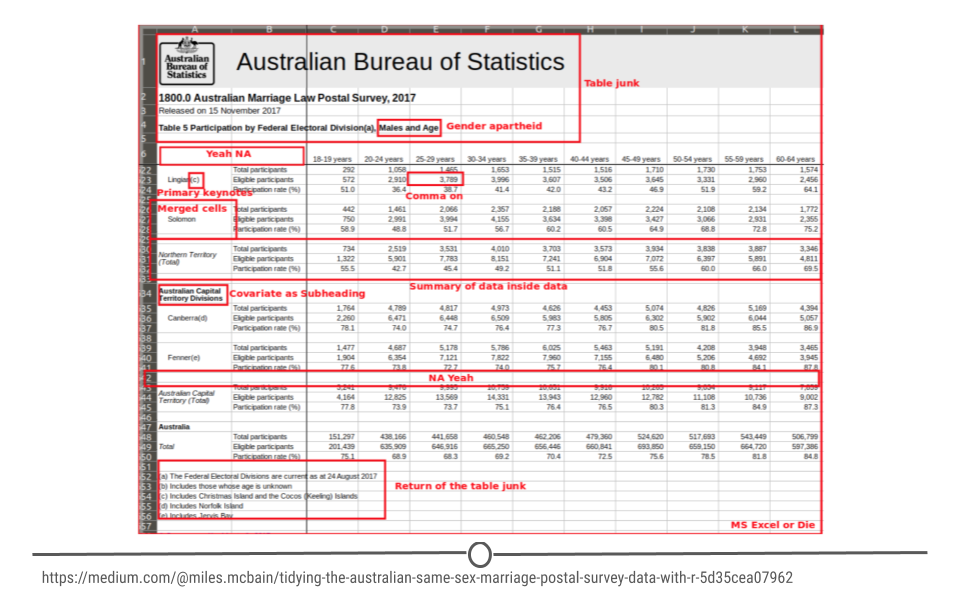
Reshaping Data
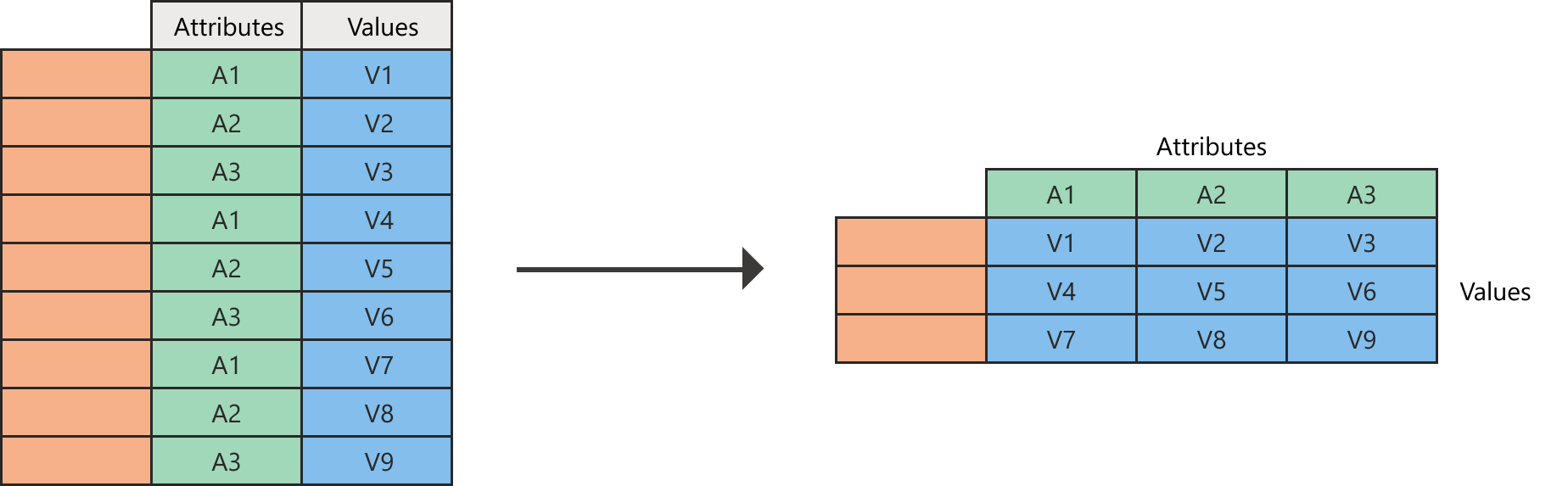
pivot_longer() — Wide to Long
Years are trapped in column names — they should be values!
pivot_longer() in Action
example_eagle_nests |>
pivot_longer(
cols = c(`2007`, `2009`),
names_to = "year",
values_to = "nests"
)# A tibble: 6 × 3
region year nests
<chr> <chr> <dbl>
1 Pacific 2007 1039
2 Pacific 2009 2587
3 Southwest 2007 51
4 Southwest 2009 176
5 Rocky Mountains and Plains 2007 200
6 Rocky Mountains and Plains 2009 338Conceptualizing pivot_longer()
When to use quotes?
"year"and"nests"are in quotes because they’re new columns- We’re creating them, not referencing existing ones
- General rule: quotes when creating, no quotes when selecting
pivot_wider() — Long to Wide
The reverse of pivot_longer():
When to Use Each?
| Direction | Function | Use case |
|---|---|---|
| Wide → Long | pivot_longer() |
Column names are data values |
| Long → Wide | pivot_wider() |
Need side-by-side comparison |
pivot_wider() in Practice
We first calculate the mean bill length by group:
Because comparing values side by side is clearer, we pivot the data wider:
Splitting Columns
separate() — Split One Column
The example_gymnastics_2 dataset has combined column names:
Step 1: Pivot Longer
example_gymnastics_2 |>
pivot_longer(
cols = -c(`country`),
names_to = "eventandyear",
values_to = "score"
)# A tibble: 12 × 3
country eventandyear score
<chr> <chr> <dbl>
1 United States vault_2012 48.1
2 United States floor_2012 45.4
3 United States vault_2016 46.9
4 United States floor_2016 46.0
5 Russia vault_2012 46.4
6 Russia floor_2012 41.6
7 Russia vault_2016 45.7
8 Russia floor_2016 42.0
9 China vault_2012 44.3
10 China floor_2012 40.8
11 China vault_2016 44.3
12 China floor_2016 42.1Step 2: Separate the Column
example_gymnastics_2 |>
pivot_longer(
cols = -c(`country`),
names_to = "eventandyear",
values_to = "score"
) |>
separate(
col = "eventandyear",
into = c("event", "year"),
sep = "_"
)# A tibble: 12 × 4
country event year score
<chr> <chr> <chr> <dbl>
1 United States vault 2012 48.1
2 United States floor 2012 45.4
3 United States vault 2016 46.9
4 United States floor 2016 46.0
5 Russia vault 2012 46.4
6 Russia floor 2012 41.6
7 Russia vault 2016 45.7
8 Russia floor 2016 42.0
9 China vault 2012 44.3
10 China floor 2012 40.8
11 China vault 2016 44.3
12 China floor 2016 42.1💡 unite() — The Reverse of separate()
unite() combines multiple columns into one:
Missing Values
drop_na() — Remove Missing Rows
# A tibble: 333 × 8
species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
<fct> <fct> <dbl> <dbl> <int> <int>
1 Adelie Torgersen 39.1 18.7 181 3750
2 Adelie Torgersen 39.5 17.4 186 3800
3 Adelie Torgersen 40.3 18 195 3250
4 Adelie Torgersen 36.7 19.3 193 3450
5 Adelie Torgersen 39.3 20.6 190 3650
6 Adelie Torgersen 38.9 17.8 181 3625
7 Adelie Torgersen 39.2 19.6 195 4675
8 Adelie Torgersen 41.1 17.6 182 3200
9 Adelie Torgersen 38.6 21.2 191 3800
10 Adelie Torgersen 34.6 21.1 198 4400
# ℹ 323 more rows
# ℹ 2 more variables: sex <fct>, year <int>drop_na() — Targeting Specific Columns
You can drop NAs in just one column:
# A tibble: 342 × 8
species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
<fct> <fct> <dbl> <dbl> <int> <int>
1 Adelie Torgersen 39.1 18.7 181 3750
2 Adelie Torgersen 39.5 17.4 186 3800
3 Adelie Torgersen 40.3 18 195 3250
4 Adelie Torgersen 36.7 19.3 193 3450
5 Adelie Torgersen 39.3 20.6 190 3650
6 Adelie Torgersen 38.9 17.8 181 3625
7 Adelie Torgersen 39.2 19.6 195 4675
8 Adelie Torgersen 34.1 18.1 193 3475
9 Adelie Torgersen 42 20.2 190 4250
10 Adelie Torgersen 37.8 17.1 186 3300
# ℹ 332 more rows
# ℹ 2 more variables: sex <fct>, year <int>Only removes rows where bill_length_mm is NA
replace_na() — Fill Missing Values
# A tibble: 344 × 8
species island bill_length_mm bill_depth_mm flipper_length_mm body_mass_g
<fct> <fct> <dbl> <dbl> <int> <int>
1 Adelie Torgersen 39.1 18.7 181 3750
2 Adelie Torgersen 39.5 17.4 186 3800
3 Adelie Torgersen 40.3 18 195 3250
4 Adelie Torgersen NA NA NA NA
5 Adelie Torgersen 36.7 19.3 193 3450
6 Adelie Torgersen 39.3 20.6 190 3650
7 Adelie Torgersen 38.9 17.8 181 3625
8 Adelie Torgersen 39.2 19.6 195 4675
9 Adelie Torgersen 34.1 18.1 193 3475
10 Adelie Torgersen 42 20.2 190 4250
# ℹ 334 more rows
# ℹ 2 more variables: sex <fct>, year <int>⚠️ Always be explicit about how you handle NAs!
A cautionary tale: bad NA handling retracted a Nature paper
Putting It All Together
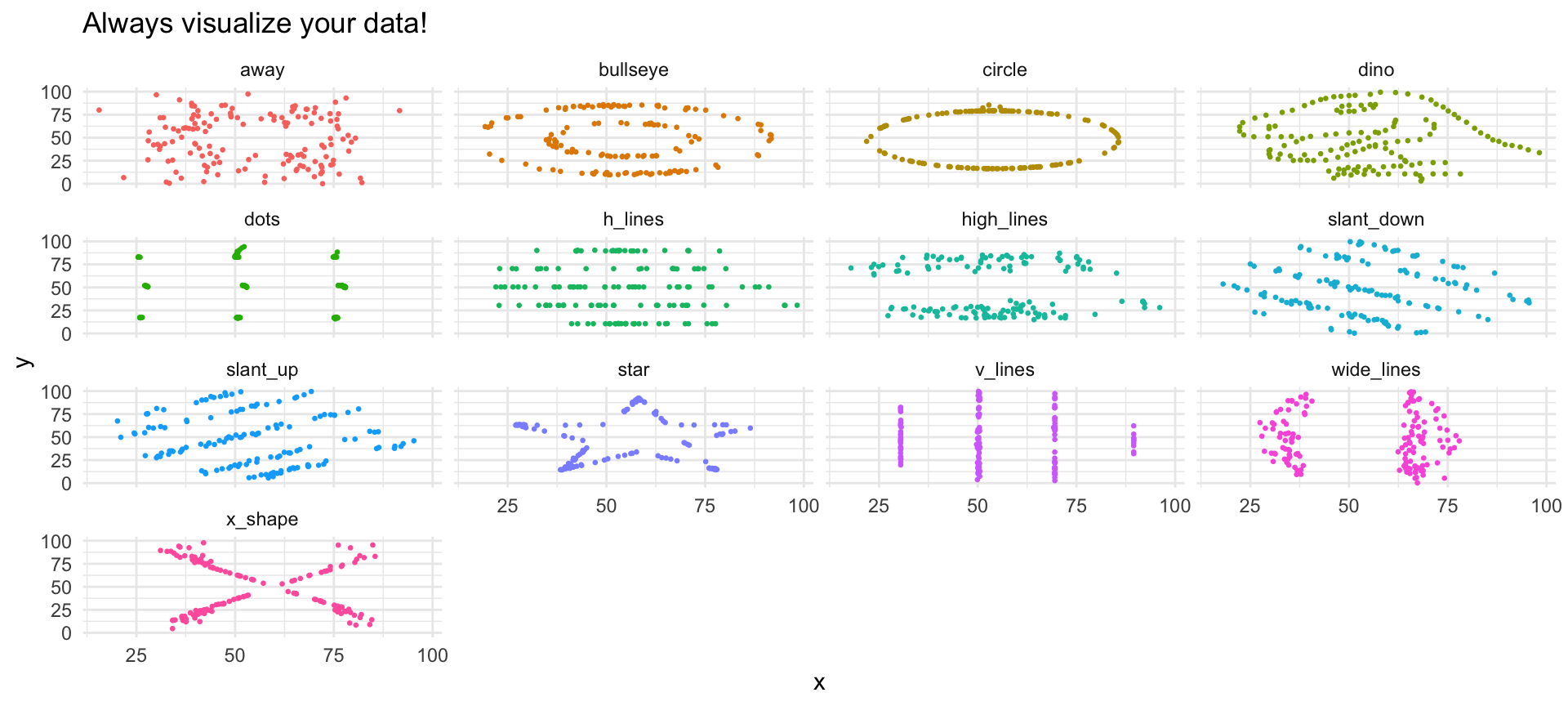
The Datasaurus Dozen 🦕
Can you trust summary statistics alone?
Let’s tidy this dataset using everything we learned today!
Step 1: Pivot Longer
Add a row ID, then pivot all columns to long format
Step 2: Separate + Pivot Wider
Split at the last underscore, then spread x/y into columns
Same Statistics…
datasaurus |>
group_by(dataset) |>
summarise(
mean_x = mean(x), mean_y = mean(y),
sd_x = sd(x), sd_y = sd(y),
cor = cor(x, y)
)# A tibble: 13 × 6
dataset mean_x mean_y sd_x sd_y cor
<chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 away 54.3 47.8 16.8 26.9 -0.0641
2 bullseye 54.3 47.8 16.8 26.9 -0.0686
3 circle 54.3 47.8 16.8 26.9 -0.0683
4 dino 54.3 47.8 16.8 26.9 -0.0645
5 dots 54.3 47.8 16.8 26.9 -0.0603
6 h_lines 54.3 47.8 16.8 26.9 -0.0617
7 high_lines 54.3 47.8 16.8 26.9 -0.0685
8 slant_down 54.3 47.8 16.8 26.9 -0.0690
9 slant_up 54.3 47.8 16.8 26.9 -0.0686
10 star 54.3 47.8 16.8 26.9 -0.0630
11 v_lines 54.3 47.8 16.8 26.9 -0.0694
12 wide_lines 54.3 47.8 16.8 26.9 -0.0666
13 x_shape 54.3 47.8 16.8 26.9 -0.0656…Completely Different Shapes!
Summary
pivot_longer()/pivot_wider()reshape dataseparate()splits,unite()combines columnsdrop_na()removes,replace_na()fills missing values- The Datasaurus shows: never trust summaries alone — always visualize!
tidyr: Tidying Data